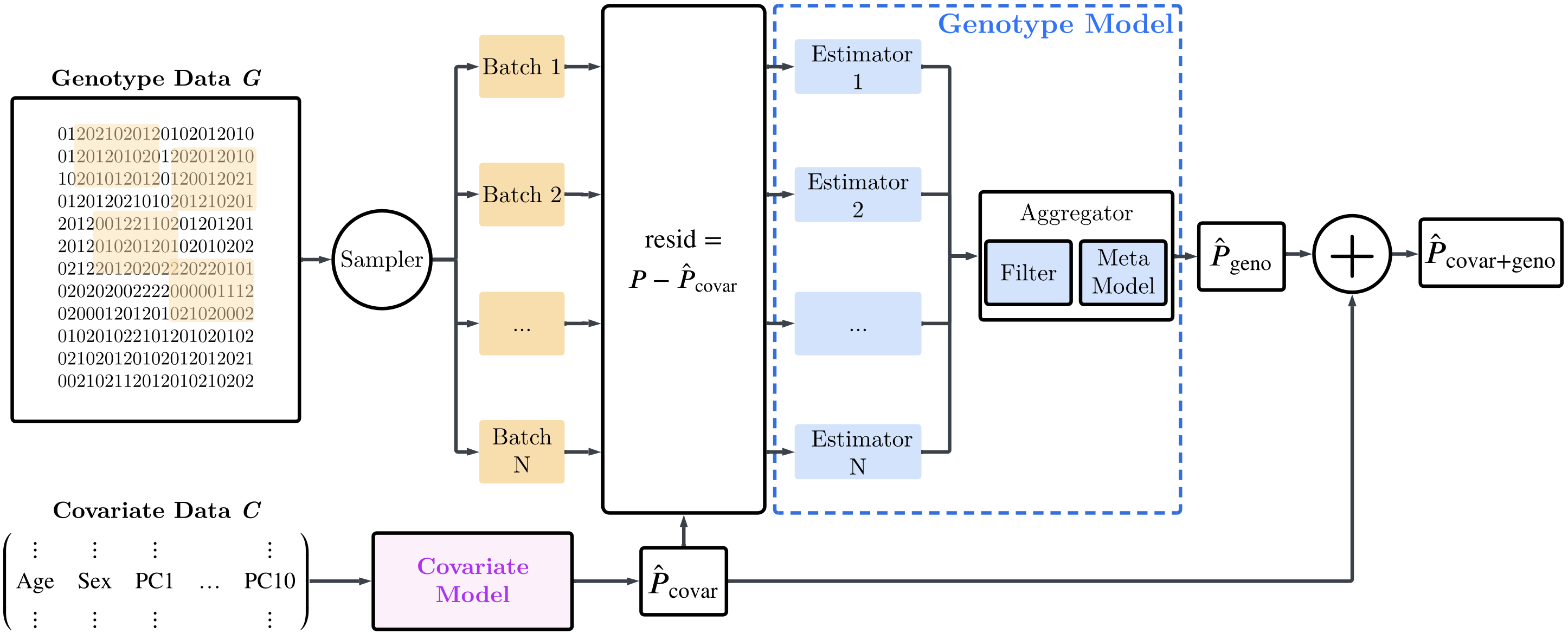
Built to model gene-gene intercations (incl. X)
GenomEn is an ensemble of (non-linear) estimators that allows to model gene-gene interactions that traditional PRS methods neglect. It natively integrates the X sex chromosome.
Inspect GenomEn's predictions and explore learned associations
We share summary statistics, predictions, variant importance scores, and intra-trait architectures derived from GenomEn models for various phenotypes. Model artifacts are also available for download.
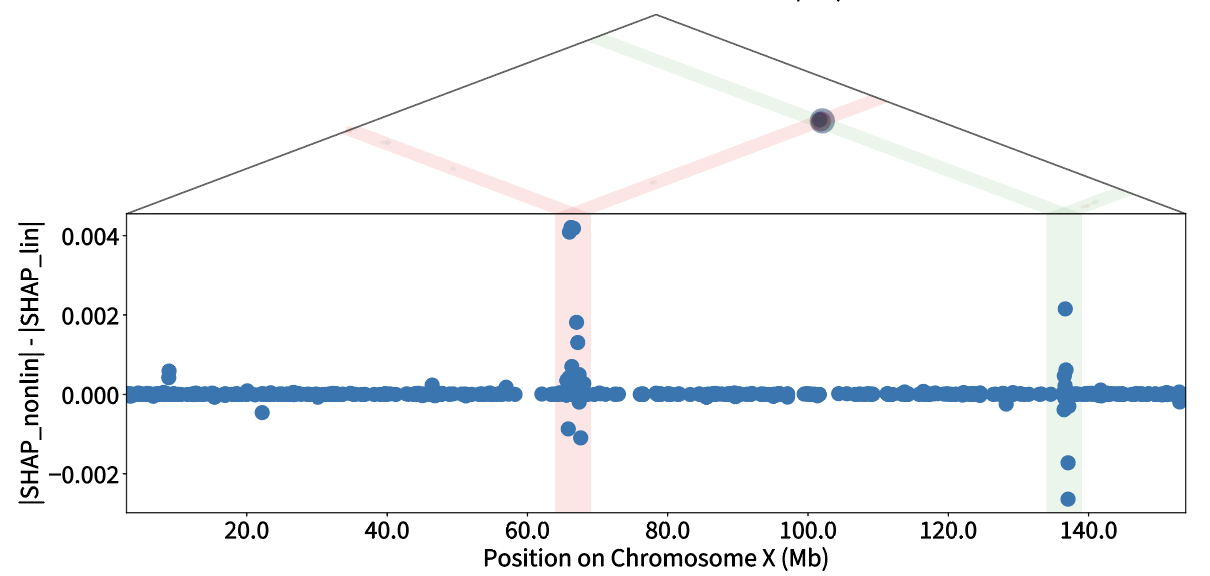
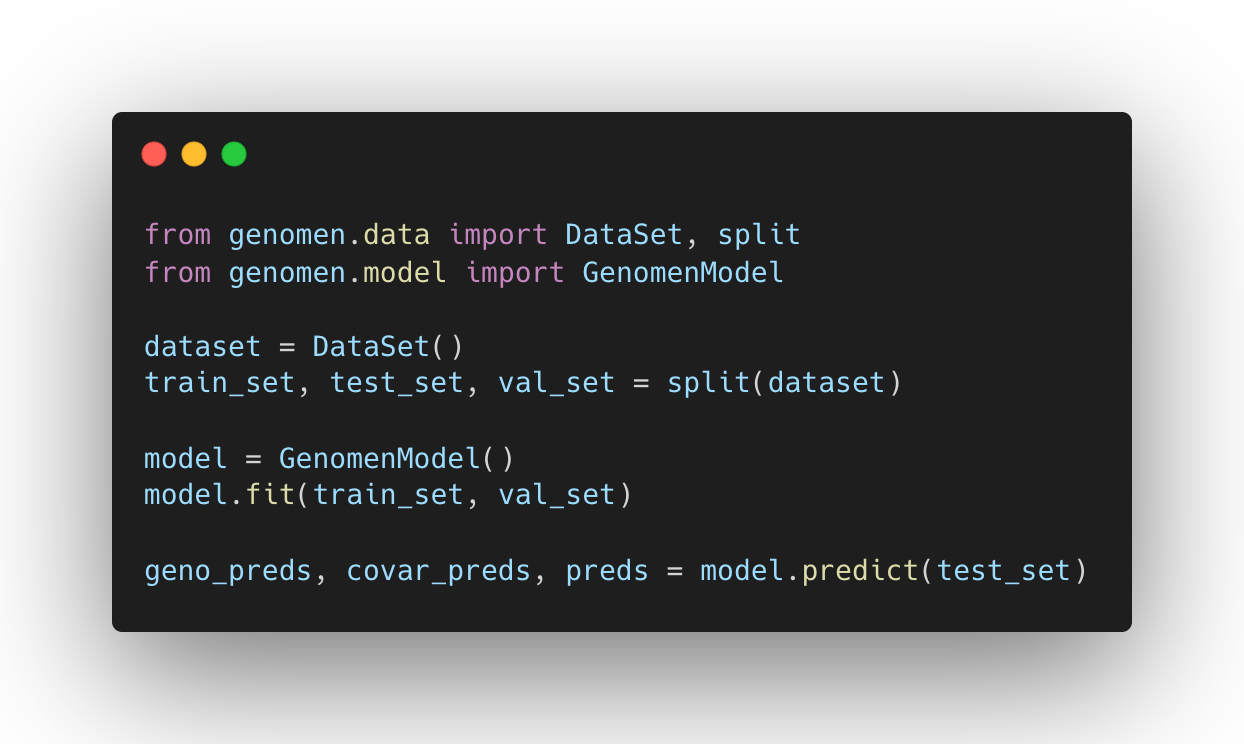
Make your own predictions with the genomen package
Use the genomen python package to train your own models, make phenotype predictions, and explore complex traits via GenomEn's variant importance values.
Explore GenomEn
BibTeX
@inproceedings{Thomassin2025GenomEn,
title = {Polygenic risk and association beyond linearity},
author = {C. Thomassin, M. Franquesa Mones, D. Bonet, P. A. Gerlach, M. Comajoan Cara, D. Mas Montserrat, and A. G. Ioannidis},
booktitle = {bioRxiv Preprint},
year = {2025},
note = {Preprint},
url = {https://your-site.com/genomen}
}