Introduction to the GenomEn package
This is a short introduction into the genomen Python package. For more in-depth information on GenomEn please refer to the paper.
We try our best to keep this tutorial up to data but if you feel like we are missing out on something, feel free to open an issue/PR on our github.
What is GenomEn?
As the name suggest GenomicEnsembling, is an ensemble of estimators used for genotype-to-phenotype prediction. The difference to PRS frameworks is that we use both lienar and non-linear estimators to also capture non-linear gene-gene interactions that are overlooked by the linear PRS methods. In our paper, we show that this allows us to improve predictive performance by an average 21.3% across 20 phenotypes compared to these traditional PRS methods. We believe this is a first step towards closing the missing heritability gap, describing the puzzling difference between theoretical heritability and predictive performance of current PRS methods to predict phenotypes from genetic material. See a brief overview of the GenomEn model architecture below.
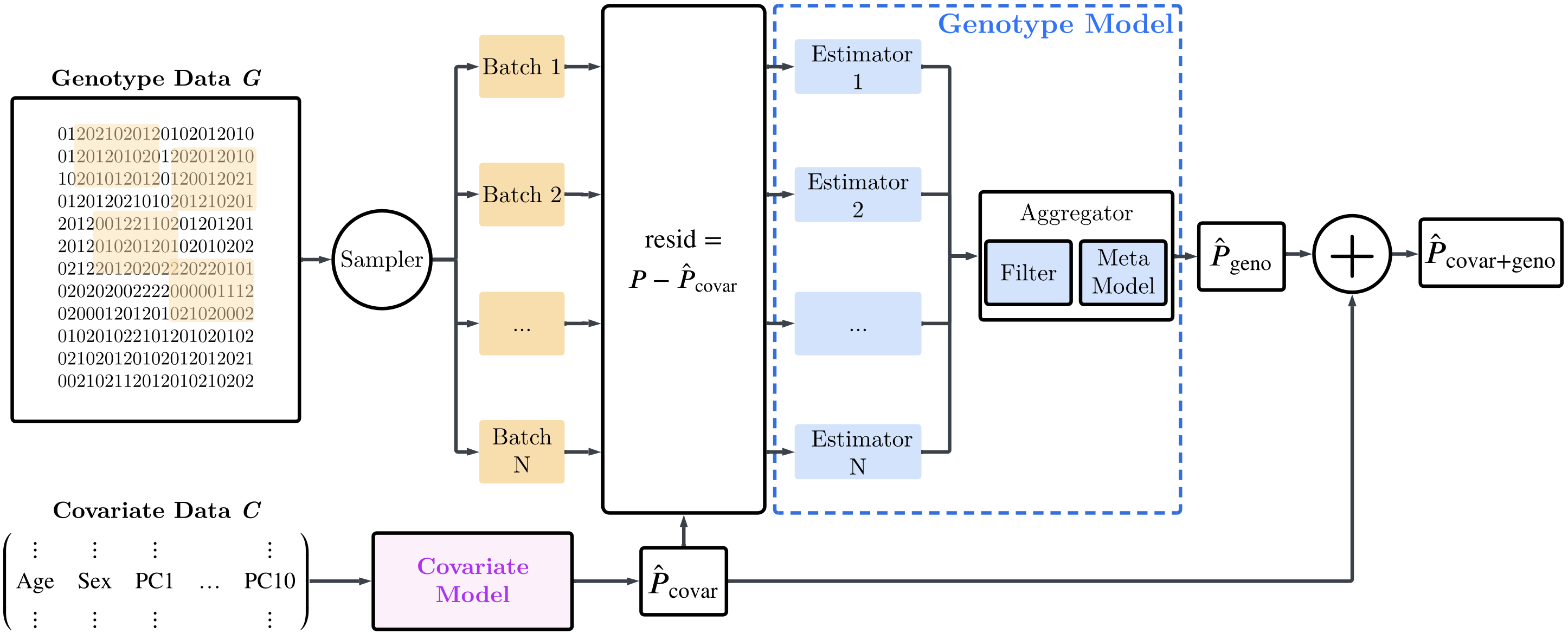
Beyond modeling of gene-gene interactions, GenomEn also natively includes variants on the X sex chromosome, variants often neglected because of difficulties in integrating them with autosomes, for phenotype prediction, further improving predictive performance and simplifying the study of X-linked traits.
The GenomEn Python package
To ensure reproducability and foster more work on closing the missing heritability gap, we publish a ready-to-use genomen Python package on PyPy for everyone to train GenomEn models on their own genetic data. The underlying code repository is public and we warmly welcome contributions and collaborations from researchers in the field and the open-source community.
We aim to make the package as user-friendly as possible by adopting scikit-learn–style APIs and training routines. Eventually, you will be able to train an entire phenotype prediction model with just a few lines as shown below.
from genomen.data import DataSet, split
from genomen.model import GenomenModel
dataset = DataSet()
train_set, test_set, val_set = split(dataset)
model = GenomenModel()
model.fit(train_set, val_set)
geno_preds, covar_preds, preds = model.predict(test_set)